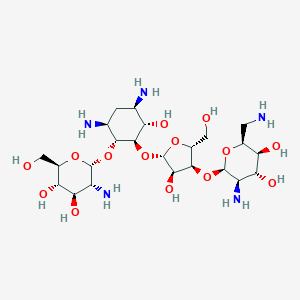
Paromomycin
Overview
Description
Paromomycin is an aminoglycoside antibiotic derived from Streptomyces rimosus var. paromomycinus, with amebicidal and antibacterial activity. This compound binds specifically to the RNA oligonucleotide at the A site of bacterial 30S ribosomes, thereby causing misreading and premature termination of translation, thereby leading to inhibition of protein synthesis followed by cell death.
This compound is an amino cyclitol glycoside that is the 1-O-(2-amino-2-deoxy-alpha-D-glucopyranoside) and the 3-O-(2,6-diamino-2,6-dideoxy-beta-L-idopyranosyl)-beta-D-ribofuranoside of 4,6-diamino-2,3-dihydroxycyclohexane (the 1R,2R,3S,4R,6S diastereoisomer). It is obtained from various Streptomyces species. A broad-spectrum antibiotic, it is used (generally as the sulfate salt) for the treatment of acute and chronic intestinal protozoal infections, but is not effective for extraintestinal protozoal infections. It is also used as a therapeutic against visceral leishmaniasis. It has a role as an antibacterial drug, an antiprotozoal drug, an anthelminthic drug and an antiparasitic agent. It is an aminoglycoside antibiotic and an amino cyclitol glycoside. It derives from a streptamine.
This compound, also known as aminosidine or estomycin, belongs to the class of organic compounds known as 4, 5-disubstituted 2-deoxystreptamines. These are 2-deoxystreptamine aminoglycosides that are glycosidically linked to a pyranose of furanose unit at the C4- and C5-positions. This compound is a drug which is used for the treatment of acute and chronic intestinal amebiasis (it is not effective in extraintestinal amebiasis). also for the management of hepatic coma as adjunctive therapy. This compound is soluble (in water) and a very weakly acidic compound (based on its pKa). This compound has been detected in multiple biofluids, such as urine and blood. Within the cell, this compound is primarily located in the cytoplasm. In humans, this compound is involved in the this compound action pathway.
Mechanism of Action
Paromomycin inhibits protein synthesis by binding to 16S ribosomal RNA. Bacterial proteins are synthesized by ribosomal RNA complexes which are composed of 2 subunits, a large subunit (50s) and small (30s) subunit, which forms a 70s ribosomal subunit. tRNA binds to the top of this ribosomal structure. Paramomycin binds to the A site, which causes defective polypeptide chains to be produced. Continuous production of defective proteins eventually leads to bacterial death.
Scientific Research Applications
Antimicrobial Activity
Paromomycin exhibits activity against both Gram-negative and Gram-positive bacteria, as well as certain protozoa and cestodes. Its mechanism involves binding to the 16S rRNA of the bacterial ribosome, leading to misreading of mRNA and subsequent inhibition of protein synthesis .
Table 1: Spectrum of Activity
| Pathogen Type | Specific Pathogens | Application |
|---|---|---|
| Bacteria | Escherichia coli, Staphylococcus aureus | Treatment of bacterial infections |
| Protozoa | Leishmania donovani, Cryptosporidium parvum | Treatment of leishmaniasis and cryptosporidiosis |
| Cestodes | Hymenolepis nana | Treatment of tapeworm infections |
Treatment of Leishmaniasis
This compound has gained prominence in the treatment of leishmaniasis, particularly visceral leishmaniasis (VL) and cutaneous leishmaniasis (CL). In India, it was licensed in 2007 for VL, demonstrating efficacy when used alone or in combination with other agents such as sodium stibogluconate .
Visceral Leishmaniasis
- Dosage : Administered at a dose of 11 mg/kg for 21 days.
- Combination Therapy : Effective when combined with sodium stibogluconate, enhancing survival rates in VL patients .
Cutaneous Leishmaniasis
- Topical Application : A combination cream of this compound and gentamicin has shown a cure rate of approximately 80% in clinical trials conducted in Tunisia and Panama .
Table 2: Clinical Trials on Leishmaniasis
| Study Location | Type | Year | Cure Rate (%) | Notes |
|---|---|---|---|---|
| Tunisia | Phase 3 trial | 2019 | ~80 | Efficacy against Leishmania major |
| Panama | Phase 3 trial | 2016 | ~80 | Compared this compound-gentamicin vs. This compound alone |
Treatment of Cryptosporidiosis
This compound is one of the few agents with demonstrated efficacy against Cryptosporidium parvum, particularly in immunocompromised individuals such as those with AIDS. Its action is effective without needing to enter the host cell cytoplasm, making it unique among treatments .
Other Applications
Beyond its use in treating parasitic infections, this compound has potential applications in research settings:
- Cell Culture Studies : Used as a reference compound for studying antileishmanial activity and RNA interactions .
- Mechanistic Studies : Investigated for its effects on protein synthesis mechanisms in various cellular models .
Case Studies
Several studies have highlighted the effectiveness of this compound in clinical settings:
-
Visceral Leishmaniasis Study :
- Conducted by the Drugs for Neglected Diseases Initiative (DNDi) in Africa.
- Showed improved survival rates when this compound was used as part of combination therapy.
-
Cutaneous Leishmaniasis Trial :
- A double-blind study comparing topical formulations demonstrated that this compound-gentamicin significantly outperformed this compound alone in terms of cure rates.
Chemical Reactions Analysis
Diazotization and Deamination
Paromomycin’s 6"'-amino group participates in diazotization reactions under acidic conditions (e.g., 40% aqueous THF at 6–7°C), yielding hydroxyl derivatives. For example:
-
6"'-Deamino-6"'-hydroxythis compound I is synthesized via diazotization of this compound, replacing the 6"'-NH₂ with a hydroxyl group. This modification retains antibacterial activity but alters ribosomal binding dynamics .
Table 1: Antibacterial Activity of this compound Derivatives
| Compound | MIC (μg/mL) S. aureus | MIC (μg/mL) E. coli |
|---|---|---|
| This compound I | 3.13 | 6.25 |
| 6"'-Deamino-6"'-hydroxythis compound I | 6.25 | 12.5 |
| Neomycin B | 1.56 | 3.13 |
| Data adapted from J-STAGE . |
Protection with Benzyloxycarbonyl (Cbz) Groups
Selective protection of amino groups is achieved using N-benzyloxycarbonyloxysuccinimide:
-
6'-N-Cbz-paromomycin and 6"'-N-Cbz-paromomycin are formed in a 31% yield mixture, which is challenging to separate. Subsequent ethoxycarbonyl (Cbe) protection of remaining amines produces pentablocked derivatives (96% yield) .
-
Catalytic hydrogenation (10% Pd/C) removes Cbz groups, regenerating free amines .
Alkylation and Acetal Formation
Modifications at the 4′,6′-positions enhance ribosomal selectivity:
-
Ethylidene acetal derivatives (e.g., compound 4 ) are synthesized via O-alkylation, improving mitochondrial ribosome targeting while retaining antibacterial efficacy .
-
Reaction of apramycin with cyclohexanecarbaldehyde dimethylacetal yields trans-dioxadecalin structures , critical for hybrid antibiotic design .
Table 2: Synthesis Outcomes of Alkylated this compound Analogues
| Derivative | Yield (%) | Key Modification |
|---|---|---|
| Ethylidene acetal 4 | 77 | 4′,6′-O-alkylidene bridge |
| 6′-Epiaprosamine 32 | 41 | 6′-Hydroxyl inversion |
| Data from PMC . |
Oxidation and Decarboxylation
Advanced synthetic routes involve oxidation followed by Barton decarboxylation:
-
This compound uronic acid (34) is generated via TEMPO-mediated oxidation of the 6′-hydroxymethyl group (79% yield) .
-
Barton decarboxylation with tert-dodecanethiol under photolysis yields 6′-deshydroxymethyl derivatives (35) in 55% yield, critical for reducing mammalian ribosome binding .
Hydrogenolysis and Deprotection
Final deprotection steps involve:
-
Palladium hydroxide-catalyzed hydrogenolysis to remove benzyloxycarbonyl groups, yielding bioactive derivatives .
-
Acidic hydrolysis (4N HCl) cleaves glycosidic bonds, releasing aprosamine and 4-aminoglucose for structural studies .
pH-Dependent Protonation Effects
This compound’s interactions with ribosomal RNA are pH-sensitive:
-
Protonation at pH < 7 enhances binding to Leishmania ribosomes via electrostatic interactions with H69 and h44 regions .
-
Structural studies confirm that protonated this compound stabilizes ribosomal subunit rotations, altering translational fidelity .
Key Findings from Experimental Data
-
Selective Activity : 6"'-hydroxyl derivatives exhibit reduced antibacterial potency compared to parent compounds but retain antileishmanial activity .
-
Hybrid Analogues : Apramycin-paromomycin hybrids (e.g., 11 ) show improved selectivity for pathogen ribosomes over mammalian counterparts .
-
Structural Dynamics : The 6′-hydroxyl group is critical for H69 interactions, as shown by crystallography (3.1 Å resolution) .
Q & A
Q. How can researchers design pharmacokinetic (PK) studies for paromomycin to account for variability in patient populations?
Answer:
- Methodological Framework: Use population pharmacokinetic modeling (e.g., nonlinear mixed-effects models) to analyze sparse and heterogeneous data from diverse cohorts, such as pediatric vs. adult patients or geographically distinct populations. Covariates like serum creatinine, albumin levels, and neutrophil counts should be included to explain interindividual variability .
- Advanced Consideration: Retrospective studies may require exclusion of physiologically implausible data points (e.g., unreliable trough samples or dosing errors), as highlighted in pooled analyses where 27.2% of this compound PK data were excluded due to variability .
Q. What methodological approaches resolve contradictions in this compound’s efficacy data across clinical trials?
Answer:
- Meta-Analysis: Apply stratified meta-analyses to compare trials with differing regimens (e.g., monotherapy vs. combination therapy) or sampling schemes. Adjust for covariates such as renal function and nutritional status (e.g., albumin levels), which influence drug exposure .
- Statistical Tools: Use Bayesian hierarchical models to account for heterogeneity in trial designs and missing data (e.g., serum creatinine data absent in Indian cohorts) .
Q. How can researchers optimize this compound dosing regimens using pharmacometric modeling?
Answer:
- Population PK/PD Modeling: Develop models integrating dose-exposure-response relationships. For example, Ethiopian trials used sparse sampling to link this compound exposure to clinical outcomes (e.g., parasite clearance), enabling dose adjustments for underweight patients .
- Covariate Analysis: Identify covariates (e.g., body weight, renal function) that significantly affect PK parameters (e.g., clearance, volume of distribution) using stepwise forward addition/backward elimination .
Q. What experimental designs address this compound’s nephrotoxicity risks in vulnerable populations?
Answer:
- Prospective Monitoring: Design trials with frequent serum creatinine and albumin measurements to detect early nephrotoxicity signals, particularly in pediatric patients (≤12 years), who constituted 59% of analyzed cohorts .
- PK/PD Safety Analysis: Use time-to-event models to correlate cumulative drug exposure with adverse outcomes (e.g., elevated creatinine), adjusting for baseline renal function .
Q. How should researchers handle missing or unreliable pharmacokinetic data in this compound studies?
Answer:
- Data Exclusion Criteria: Predefine exclusion thresholds (e.g., trough samples collected post-next dose, quantification errors) to maintain data integrity. In one study, 3/232 this compound observations were excluded due to sampling errors .
- Imputation Methods: Apply multiple imputation or full PK modeling (e.g., maximum likelihood estimation) to handle missing covariates like serum creatinine in retrospective datasets .
Q. What advanced statistical methods validate this compound’s therapeutic equivalence in combination therapies?
Answer:
- Non-Inferiority Trials: Use Bayesian adaptive designs to compare this compound-miltefosine combination therapy against standard regimens. Analyze PK interactions (e.g., drug-drug effects on clearance) through joint modeling .
- Contradiction Resolution: Apply causal inference methods (e.g., propensity score matching) to adjust for confounding variables in observational studies where treatment groups are imbalanced .
Q. How can researchers ensure reproducibility in this compound’s preclinical efficacy models?
Answer:
- Standardized Protocols: Adopt consistent in vitro assays (e.g., minimum inhibitory concentration [MIC] testing) and animal models (e.g., Leishmania-infected rodents) with documented quality controls .
- Data Transparency: Publish raw pharmacokinetic data, exclusion criteria, and model code in repositories like Dryad or Figshare to enable independent validation .
Q. What strategies mitigate bias in multicenter this compound trials with heterogeneous protocols?
Answer:
- Centralized Monitoring: Implement real-time data audits to standardize sampling times and dosing records across sites, as done in Ethiopian and Indian trials .
- Covariate Adjustment: Use mixed-effects models to account for site-specific variability (e.g., differences in nutritional status or healthcare infrastructure) .
How can researchers formulate PICOT-compliant questions for this compound studies?
Answer:
- PICOT Framework:
- P: Patients with visceral leishmaniasis (e.g., pediatric vs. adult).
- I: this compound 15 mg/kg/day intramuscularly.
- C: Miltefosine monotherapy.
- O: Parasite clearance at Day 28.
- T: 28-day follow-up.
- Advanced Application: Use FINER criteria (Feasible, Interesting, Novel, Ethical, Relevant) to evaluate question rigor, ensuring alignment with gaps in PK/PD understanding .
Q. What methodologies reconcile contradictory findings in this compound’s resistance mechanisms?
Answer:
- Genomic Analysis: Perform whole-genome sequencing of resistant Leishmania strains to identify mutations (e.g., ribosomal RNA gene alterations) linked to reduced drug binding .
- In Vitro/In Vivo Correlation: Use hollow-fiber infection models to simulate human PK profiles and validate resistance thresholds observed in clinical isolates .
Properties
IUPAC Name |
(2S,3S,4R,5R,6R)-5-amino-2-(aminomethyl)-6-[(2R,3S,4R,5S)-5-[(1R,2R,3S,5R,6S)-3,5-diamino-2-[(2S,3R,4R,5S,6R)-3-amino-4,5-dihydroxy-6-(hydroxymethyl)oxan-2-yl]oxy-6-hydroxycyclohexyl]oxy-4-hydroxy-2-(hydroxymethyl)oxolan-3-yl]oxyoxane-3,4-diol | |
|---|---|---|
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
InChI |
InChI=1S/C23H45N5O14/c24-2-7-13(32)15(34)10(27)21(37-7)41-19-9(4-30)39-23(17(19)36)42-20-12(31)5(25)1-6(26)18(20)40-22-11(28)16(35)14(33)8(3-29)38-22/h5-23,29-36H,1-4,24-28H2/t5-,6+,7+,8-,9-,10-,11-,12+,13-,14-,15-,16-,17-,18-,19-,20-,21-,22-,23+/m1/s1 | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
InChI Key |
UOZODPSAJZTQNH-LSWIJEOBSA-N | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Canonical SMILES |
C1C(C(C(C(C1N)OC2C(C(C(C(O2)CO)O)O)N)OC3C(C(C(O3)CO)OC4C(C(C(C(O4)CN)O)O)N)O)O)N | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Isomeric SMILES |
C1[C@H]([C@@H]([C@H]([C@@H]([C@H]1N)O[C@@H]2[C@@H]([C@H]([C@@H]([C@H](O2)CO)O)O)N)O[C@H]3[C@@H]([C@@H]([C@H](O3)CO)O[C@@H]4[C@@H]([C@H]([C@@H]([C@@H](O4)CN)O)O)N)O)O)N | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Molecular Formula |
C23H45N5O14 | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Related CAS |
1263-89-4 (sulfate), 35665-49-7 (sulfate (2:5)), 7205-49-4 (sulfate (1:1)) | |
| Record name | Paromomycin [INN:BAN] | |
| Source | ChemIDplus | |
| URL | https://pubchem.ncbi.nlm.nih.gov/substance/?source=chemidplus&sourceid=0007542372 | |
| Description | ChemIDplus is a free, web search system that provides access to the structure and nomenclature authority files used for the identification of chemical substances cited in National Library of Medicine (NLM) databases, including the TOXNET system. | |
DSSTOX Substance ID |
DTXSID8023424 | |
| Record name | Paromomycin | |
| Source | EPA DSSTox | |
| URL | https://comptox.epa.gov/dashboard/DTXSID8023424 | |
| Description | DSSTox provides a high quality public chemistry resource for supporting improved predictive toxicology. | |
Molecular Weight |
615.6 g/mol | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Physical Description |
Solid | |
| Record name | Paromomycin | |
| Source | Human Metabolome Database (HMDB) | |
| URL | http://www.hmdb.ca/metabolites/HMDB0015490 | |
| Description | The Human Metabolome Database (HMDB) is a freely available electronic database containing detailed information about small molecule metabolites found in the human body. | |
| Explanation | HMDB is offered to the public as a freely available resource. Use and re-distribution of the data, in whole or in part, for commercial purposes requires explicit permission of the authors and explicit acknowledgment of the source material (HMDB) and the original publication (see the HMDB citing page). We ask that users who download significant portions of the database cite the HMDB paper in any resulting publications. | |
Solubility |
7.97e+01 g/L | |
| Record name | Paromomycin | |
| Source | Human Metabolome Database (HMDB) | |
| URL | http://www.hmdb.ca/metabolites/HMDB0015490 | |
| Description | The Human Metabolome Database (HMDB) is a freely available electronic database containing detailed information about small molecule metabolites found in the human body. | |
| Explanation | HMDB is offered to the public as a freely available resource. Use and re-distribution of the data, in whole or in part, for commercial purposes requires explicit permission of the authors and explicit acknowledgment of the source material (HMDB) and the original publication (see the HMDB citing page). We ask that users who download significant portions of the database cite the HMDB paper in any resulting publications. | |
Mechanism of Action |
Paromomycin inhibits protein synthesis by binding to 16S ribosomal RNA. Bacterial proteins are synthesized by ribosomal RNA complexes which are composed of 2 subunits, a large subunit (50s) and small (30s) subunit, which forms a 70s ribosomal subunit. tRNA binds to the top of this ribosomal structure. Paramomycin binds to the A site, which causes defective polypeptide chains to be produced. Continuous production of defective proteins eventually leads to bacterial death. | |
| Record name | Paromomycin | |
| Source | DrugBank | |
| URL | https://www.drugbank.ca/drugs/DB01421 | |
| Description | The DrugBank database is a unique bioinformatics and cheminformatics resource that combines detailed drug (i.e. chemical, pharmacological and pharmaceutical) data with comprehensive drug target (i.e. sequence, structure, and pathway) information. | |
| Explanation | Creative Common's Attribution-NonCommercial 4.0 International License (http://creativecommons.org/licenses/by-nc/4.0/legalcode) | |
CAS No. |
7542-37-2, 1263-89-4 | |
| Record name | Paromomycin | |
| Source | CAS Common Chemistry | |
| URL | https://commonchemistry.cas.org/detail?cas_rn=7542-37-2 | |
| Description | CAS Common Chemistry is an open community resource for accessing chemical information. Nearly 500,000 chemical substances from CAS REGISTRY cover areas of community interest, including common and frequently regulated chemicals, and those relevant to high school and undergraduate chemistry classes. This chemical information, curated by our expert scientists, is provided in alignment with our mission as a division of the American Chemical Society. | |
| Explanation | The data from CAS Common Chemistry is provided under a CC-BY-NC 4.0 license, unless otherwise stated. | |
| Record name | Paromomycin [INN:BAN] | |
| Source | ChemIDplus | |
| URL | https://pubchem.ncbi.nlm.nih.gov/substance/?source=chemidplus&sourceid=0007542372 | |
| Description | ChemIDplus is a free, web search system that provides access to the structure and nomenclature authority files used for the identification of chemical substances cited in National Library of Medicine (NLM) databases, including the TOXNET system. | |
| Record name | Paromomycin | |
| Source | DrugBank | |
| URL | https://www.drugbank.ca/drugs/DB01421 | |
| Description | The DrugBank database is a unique bioinformatics and cheminformatics resource that combines detailed drug (i.e. chemical, pharmacological and pharmaceutical) data with comprehensive drug target (i.e. sequence, structure, and pathway) information. | |
| Explanation | Creative Common's Attribution-NonCommercial 4.0 International License (http://creativecommons.org/licenses/by-nc/4.0/legalcode) | |
| Record name | Paromomycin | |
| Source | EPA DSSTox | |
| URL | https://comptox.epa.gov/dashboard/DTXSID8023424 | |
| Description | DSSTox provides a high quality public chemistry resource for supporting improved predictive toxicology. | |
| Record name | Paromomycin | |
| Source | European Chemicals Agency (ECHA) | |
| URL | https://echa.europa.eu/substance-information/-/substanceinfo/100.028.567 | |
| Description | The European Chemicals Agency (ECHA) is an agency of the European Union which is the driving force among regulatory authorities in implementing the EU's groundbreaking chemicals legislation for the benefit of human health and the environment as well as for innovation and competitiveness. | |
| Explanation | Use of the information, documents and data from the ECHA website is subject to the terms and conditions of this Legal Notice, and subject to other binding limitations provided for under applicable law, the information, documents and data made available on the ECHA website may be reproduced, distributed and/or used, totally or in part, for non-commercial purposes provided that ECHA is acknowledged as the source: "Source: European Chemicals Agency, http://echa.europa.eu/". Such acknowledgement must be included in each copy of the material. ECHA permits and encourages organisations and individuals to create links to the ECHA website under the following cumulative conditions: Links can only be made to webpages that provide a link to the Legal Notice page. | |
| Record name | PAROMOMYCIN | |
| Source | FDA Global Substance Registration System (GSRS) | |
| URL | https://gsrs.ncats.nih.gov/ginas/app/beta/substances/61JJC8N5ZK | |
| Description | The FDA Global Substance Registration System (GSRS) enables the efficient and accurate exchange of information on what substances are in regulated products. Instead of relying on names, which vary across regulatory domains, countries, and regions, the GSRS knowledge base makes it possible for substances to be defined by standardized, scientific descriptions. | |
| Explanation | Unless otherwise noted, the contents of the FDA website (www.fda.gov), both text and graphics, are not copyrighted. They are in the public domain and may be republished, reprinted and otherwise used freely by anyone without the need to obtain permission from FDA. Credit to the U.S. Food and Drug Administration as the source is appreciated but not required. | |
| Record name | Paromomycin | |
| Source | Human Metabolome Database (HMDB) | |
| URL | http://www.hmdb.ca/metabolites/HMDB0015490 | |
| Description | The Human Metabolome Database (HMDB) is a freely available electronic database containing detailed information about small molecule metabolites found in the human body. | |
| Explanation | HMDB is offered to the public as a freely available resource. Use and re-distribution of the data, in whole or in part, for commercial purposes requires explicit permission of the authors and explicit acknowledgment of the source material (HMDB) and the original publication (see the HMDB citing page). We ask that users who download significant portions of the database cite the HMDB paper in any resulting publications. | |
Disclaimer and Information on In-Vitro Research Products
Please be aware that all articles and product information presented on BenchChem are intended solely for informational purposes. The products available for purchase on BenchChem are specifically designed for in-vitro studies, which are conducted outside of living organisms. In-vitro studies, derived from the Latin term "in glass," involve experiments performed in controlled laboratory settings using cells or tissues. It is important to note that these products are not categorized as medicines or drugs, and they have not received approval from the FDA for the prevention, treatment, or cure of any medical condition, ailment, or disease. We must emphasize that any form of bodily introduction of these products into humans or animals is strictly prohibited by law. It is essential to adhere to these guidelines to ensure compliance with legal and ethical standards in research and experimentation.