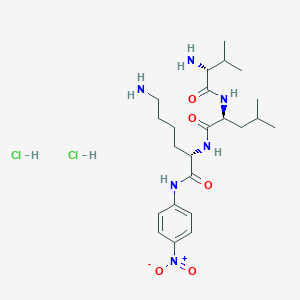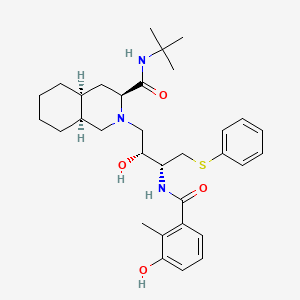
Nelfinavir
Overview
Description
Nelfinavir is an antiretroviral medication used in the treatment of HIV/AIDS. This compound is an orally bioavailable human immunodeficiency virus HIV-1 protease inhibitor and was patented in 1992 and approved for medical use in 1997 .
Preparation Methods
Nelfinavir can be synthesized through various methods. One approach involves the development of nanocrystals of this compound Mesylate to overcome the drawbacks associated with the drug, such as poor solubility and oral bioavailability . The nanocrystals are prepared using a combination technique and ultrasonication method with polyvinyl alcohol and poloxamer 407 as stabilizers . The solid-state characteristics of the optimized nanocrystals are studied using X-ray diffraction, Fourier-transform infrared spectroscopy, differential scanning calorimetry, and scanning electron microscopy .
Chemical Reactions Analysis
Nelfinavir undergoes various chemical reactions, including oxidation, reduction, and substitution. It is a protease inhibitor with activity against Human Immunodeficiency Virus Type 1 (HIV-1) . Protease inhibitors block the part of HIV called protease, which is an enzyme required for the proteolytic cleavage of the viral polyprotein precursors into the individual functional proteins found in infectious HIV-1 . This compound binds to the protease active site and inhibits the activity of the enzyme, preventing the cleavage of the viral polyproteins and resulting in the formation of immature non-infectious viral particles .
Scientific Research Applications
Antiviral Applications
HIV Treatment
Nelfinavir is primarily known as an HIV protease inhibitor, playing a crucial role in highly active antiretroviral therapy (HAART). It works by inhibiting the HIV protease enzyme, which is essential for the maturation of infectious viral particles. This action effectively reduces viral load in patients and improves immune function.
Oncological Applications
This compound has demonstrated significant antitumor effects across various cancer types through multiple mechanisms:
2.1 Mechanisms of Action
- Inhibition of Signaling Pathways : this compound suppresses the Akt signaling pathway, which is often dysregulated in cancer cells. By inhibiting this pathway, this compound induces apoptosis and impairs cell proliferation in several cancer types, including small-cell lung cancer (SCLC) and castration-resistant prostate cancer (CRPC) .
- Induction of Unfolded Protein Response (UPR) : In SCLC cells, this compound triggers UPR, leading to cell death. This response is linked to the inhibition of mTOR activation and reduced expression of oncogenic markers like achaete-scute homolog 1 (ASCL1) .
2.2 Case Studies and Clinical Trials
- Small-Cell Lung Cancer : In vitro studies have shown that this compound effectively inhibits SCLC cell proliferation and induces apoptosis at concentrations as low as 10 μM . In vivo experiments with patient-derived xenograft (PDX) models confirmed its efficacy in reducing tumor growth.
- Castration-Resistant Prostate Cancer : this compound has been identified as a potent inhibitor of S2P cleavage, a process vital for CRPC proliferation. This inhibition leads to increased ER stress and accumulation of inactive precursors of key transcription factors . Clinical trials are being proposed to further explore its therapeutic potential in this context.
Repurposing for Other Diseases
Recent studies suggest that this compound may be effective against other infectious diseases:
- Helminth Infections : A study indicated that this compound shows promise in treating echinococcosis caused by Echinococcus multilocularis. Its repurposing could provide a single-drug therapy option for patients co-infected with HIV and helminths .
Pharmacokinetics and Safety Profile
This compound's pharmacokinetics have been extensively studied to optimize dosing regimens in HIV-infected populations. Variability in drug metabolism can influence therapeutic outcomes, necessitating careful monitoring during treatment .
Summary Table of this compound Applications
Mechanism of Action
Nelfinavir exerts its effects by inhibiting the HIV viral proteinase enzyme, which prevents the cleavage of the gag-pol polyprotein, resulting in noninfectious, immature viral particles . The inhibition of the protease enzyme is essential for the HIV life cycle, as it prevents the assembly of immature virus proteins into mature, infectious virions . This compound binds to the protease active site and inhibits its activity, leading to the formation of immature non-infectious viral particles .
Comparison with Similar Compounds
Nelfinavir is unique among protease inhibitors due to its specific structure and mechanism of action. Similar compounds include other protease inhibitors such as ritonavir, lopinavir, and indinavir . These compounds also inhibit the HIV protease enzyme but may differ in their pharmacokinetic properties, side effects, and efficacy . This compound has been found to have a unique cis-decahydroisoquinoline-2-carboxamide moiety, which may provide the structural basis for its increased efficacy against cancer compared to other HIV protease inhibitors .
Biological Activity
Nelfinavir, an HIV protease inhibitor, has garnered attention not only for its antiviral properties but also for its potential anticancer activity. This article explores the biological activity of this compound, detailing its mechanisms of action, clinical applications, and research findings.
This compound exhibits multiple mechanisms that contribute to its biological activity:
- Proteasome Inhibition : this compound inhibits proteasome activity, leading to the accumulation of misfolded proteins and inducing endoplasmic reticulum (ER) stress. This process triggers the unfolded protein response (UPR), which is crucial for cellular homeostasis and can lead to apoptosis in cancer cells .
- Disruption of Lipid Membranes : Research indicates that this compound interacts with lipid bilayers in cellular organelles, causing lipid bilayer stress. This disruption affects mitochondrial respiration and transmembrane protein transport, contributing to its anticancer effects .
- Inhibition of Signaling Pathways : this compound has been shown to suppress the Akt signaling pathway, which is vital for cell survival and metabolism. It also inhibits multiple members of the protein kinase-like superfamily, impacting various biological processes associated with carcinogenesis and metastasis .
Clinical Applications
This compound's anticancer potential has been evaluated in several clinical settings:
- Multiple Myeloma : In a phase I trial (SAKK 65/08), this compound demonstrated significant activity against multiple myeloma (MM), particularly in patients who were refractory to proteasome inhibitors. The combination of this compound with bortezomib and dexamethasone yielded an overall response rate of 65% in heavily pretreated patients .
- Broad Anticancer Activity : this compound has shown broad anticancer activity across various preclinical models. Its ability to induce apoptosis and cell cycle arrest positions it as a promising candidate for further investigation in cancer therapy .
Research Findings
Recent studies have elucidated the biological activity of this compound through various experimental approaches:
Table 1: Summary of Key Studies on this compound
Case Studies
-
Case Study: this compound in Multiple Myeloma
- A patient with advanced MM received this compound as part of a combination therapy. The treatment resulted in a partial response, highlighting this compound's potential as an effective chemotherapeutic agent in this setting.
-
Case Study: Anticancer Mechanisms
- In vitro studies demonstrated that this compound could induce apoptosis in various cancer cell lines by activating ER stress pathways and disrupting mitochondrial function, suggesting a multifaceted approach to cancer treatment.
Q & A
Basic Research Questions
Q. What experimental models are commonly used to evaluate nelfinavir's anticancer efficacy, and how do they inform mechanistic insights?
- Methodological Answer : Preclinical studies often use in vitro cancer cell lines (e.g., SCLC, breast cancer) and in vivo patient-derived xenograft (PDX) models. For example:
- Cell viability assays (MTT, flow cytometry) assess apoptosis via Annexin V/PI staining (e.g., MDA-MB231, MCF-7) .
- Western blotting evaluates biomarkers like Bak, cytochrome c, and caspases to confirm apoptosis .
- PDX models validate tumor regression and molecular pathways (e.g., mTOR inhibition, UPR induction) in SCLC .
Q. How is this compound quantified in pharmacokinetic studies, and what analytical methods ensure accuracy?
- Methodological Answer : UV-Visible spectrophotometry (e.g., λ = 210–310 nm) with validation parameters:
- Linearity : 10–60 µg/ml for this compound (R² > 0.999) .
- Calibration : Equations like are derived via least squares regression .
LC-MS/MS is preferred for plasma metabolite quantification (e.g., M8 levels: 0.55–1.96 µM) .
Q. What is the role of this compound's active metabolite M8 in antiviral and anticancer activity?
- Methodological Answer :
- CYP2C19 metabolism : Generates M8, which retains anti-HIV activity (EC₅₀: 34 nM vs. This compound’s 30 nM) and binds PXR competitively .
- Docking simulations : M8 shows distinct binding poses in PXR’s ligand-binding pocket (LBP), influencing receptor antagonism .
Advanced Research Questions
Q. How does this compound modulate oxidative stress to induce cancer cell death, and what experimental approaches validate this?
- Methodological Answer :
- ROS assays : H₂DCF-DA staining quantifies ROS accumulation (e.g., 2–3-fold increase in MDA-MB231 cells) .
- Lipid peroxidation : MDA levels (nM) correlate with this compound-induced oxidative damage .
- Akt pathway disruption : ROS-dependent degradation of Akt-HSP90 complexes, validated via immunoprecipitation .
Q. What computational methods predict this compound’s off-target kinase interactions, and how do they align with experimental data?
- Methodological Answer :
- MM/GBSA free energy calculations : Predict binding affinities for EGFR (ΔG = −50 kcal/mol) but not FGFR/EPHB4 .
- Molecular docking : Identifies ATP-binding site interactions (e.g., EGFR vs. lapatinib) .
Discrepancies arise for Akt2/CDK2, where computational predictions lack experimental validation .
Q. How do pharmacogenomic variants influence this compound’s plasma exposure and clinical outcomes?
- Methodological Answer :
- CYP2B6 516G→T : Correlates with 30% higher efavirenz exposure (p < 0.01) .
- CYP2C19 681G→A : Reduces this compound clearance, linked to 40% lower virologic failure risk (p = 0.039) .
- MDR1 3435 TT genotype : Associated with reduced resistance emergence in HIV trials (OR = 0.6) .
Q. What mechanisms underlie this compound’s radiosensitizing effects, and how are they tested in combination therapies?
- Methodological Answer :
- HIF-1α/VEGF suppression : Western blotting shows this compound reduces HIF-1α by 70% under hypoxia, validated via EF5 hypoxia marker assays .
- In vivo xenografts : Tumor regrowth delay (2–3 weeks) with this compound + radiation vs. radiation alone .
- Synergy with cisplatin : PPP2R1A silencing enhances apoptosis (p < 0.001) in LUAD models .
Q. Methodological Considerations
Q. How are contradictions in this compound’s efficacy across cancer cell lines addressed experimentally?
- Answer :
- Cell line stratification : Compare genetic profiles (e.g., H69 SCLC cells with PIK3CA mutations resist mTOR inhibition) .
- Biomarker panels : Use ATF4, CHOP, and SESN2 to differentiate UPR vs. mTOR-driven responses .
- Pharmacological manipulation : Co-treatment with rapamycin/tunicamycin isolates pathway-specific effects .
Q. What in silico and in vitro assays resolve this compound’s nuclear receptor agonism/antagonism?
- Answer :
- Luciferase reporter assays : HepG2 cells transfected with PXR/CAR show partial agonism (2-fold induction) and competitive antagonism (IC₅₀ = 10 µM) .
- Limited proteolysis : PXR-LBD structural changes confirm ligand binding (30–250 µM this compound) .
- AlphaSphere analysis : Identifies alternative binding sites (e.g., 1M13, 2O9I) via docking simulations .
Properties
IUPAC Name |
(3S,4aS,8aS)-N-tert-butyl-2-[(2R,3R)-2-hydroxy-3-[(3-hydroxy-2-methylbenzoyl)amino]-4-phenylsulfanylbutyl]-3,4,4a,5,6,7,8,8a-octahydro-1H-isoquinoline-3-carboxamide | |
|---|---|---|
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
InChI |
InChI=1S/C32H45N3O4S/c1-21-25(15-10-16-28(21)36)30(38)33-26(20-40-24-13-6-5-7-14-24)29(37)19-35-18-23-12-9-8-11-22(23)17-27(35)31(39)34-32(2,3)4/h5-7,10,13-16,22-23,26-27,29,36-37H,8-9,11-12,17-20H2,1-4H3,(H,33,38)(H,34,39)/t22-,23+,26-,27-,29+/m0/s1 | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
InChI Key |
QAGYKUNXZHXKMR-HKWSIXNMSA-N | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Canonical SMILES |
CC1=C(C=CC=C1O)C(=O)NC(CSC2=CC=CC=C2)C(CN3CC4CCCCC4CC3C(=O)NC(C)(C)C)O | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Isomeric SMILES |
CC1=C(C=CC=C1O)C(=O)N[C@@H](CSC2=CC=CC=C2)[C@@H](CN3C[C@H]4CCCC[C@H]4C[C@H]3C(=O)NC(C)(C)C)O | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Molecular Formula |
C32H45N3O4S | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Related CAS |
159989-65-8 (monomethane sulfonate (salt)) | |
| Record name | Nelfinavir [INN:BAN] | |
| Source | ChemIDplus | |
| URL | https://pubchem.ncbi.nlm.nih.gov/substance/?source=chemidplus&sourceid=0159989647 | |
| Description | ChemIDplus is a free, web search system that provides access to the structure and nomenclature authority files used for the identification of chemical substances cited in National Library of Medicine (NLM) databases, including the TOXNET system. | |
DSSTOX Substance ID |
DTXSID5035080 | |
| Record name | Nelfinavir | |
| Source | EPA DSSTox | |
| URL | https://comptox.epa.gov/dashboard/DTXSID5035080 | |
| Description | DSSTox provides a high quality public chemistry resource for supporting improved predictive toxicology. | |
Molecular Weight |
567.8 g/mol | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Physical Description |
Solid | |
| Record name | Nelfinavir | |
| Source | Human Metabolome Database (HMDB) | |
| URL | http://www.hmdb.ca/metabolites/HMDB0014365 | |
| Description | The Human Metabolome Database (HMDB) is a freely available electronic database containing detailed information about small molecule metabolites found in the human body. | |
| Explanation | HMDB is offered to the public as a freely available resource. Use and re-distribution of the data, in whole or in part, for commercial purposes requires explicit permission of the authors and explicit acknowledgment of the source material (HMDB) and the original publication (see the HMDB citing page). We ask that users who download significant portions of the database cite the HMDB paper in any resulting publications. | |
Solubility |
Slightly soluble, 1.91e-03 g/L | |
| Record name | Nelfinavir | |
| Source | DrugBank | |
| URL | https://www.drugbank.ca/drugs/DB00220 | |
| Description | The DrugBank database is a unique bioinformatics and cheminformatics resource that combines detailed drug (i.e. chemical, pharmacological and pharmaceutical) data with comprehensive drug target (i.e. sequence, structure, and pathway) information. | |
| Explanation | Creative Common's Attribution-NonCommercial 4.0 International License (http://creativecommons.org/licenses/by-nc/4.0/legalcode) | |
| Record name | Nelfinavir | |
| Source | Human Metabolome Database (HMDB) | |
| URL | http://www.hmdb.ca/metabolites/HMDB0014365 | |
| Description | The Human Metabolome Database (HMDB) is a freely available electronic database containing detailed information about small molecule metabolites found in the human body. | |
| Explanation | HMDB is offered to the public as a freely available resource. Use and re-distribution of the data, in whole or in part, for commercial purposes requires explicit permission of the authors and explicit acknowledgment of the source material (HMDB) and the original publication (see the HMDB citing page). We ask that users who download significant portions of the database cite the HMDB paper in any resulting publications. | |
Mechanism of Action |
HIV viral protease is an important enzyme for HIV maturation and pathogenicity since HIV produces its structural and key proteins in the form of a polyprotein that needs to be cleaved by a protease. HIV protease is synthesized as part of the Gag-pol polyprotein, where Gag encodes for the capsid and matrix protein to form the outer protein shell, and Pol encodes for the reverse transcriptase and integrase protein to synthesize and incorporate its genome into host cells. The Gag-pol polyprotein undergoes proteolytic cleavage by HIV protease to produce 66 molecular species which will assume conformational changes to become fully active. Inhibition of protease, therefore, prevents HIV virion from fully maturing and becoming infective. Nelfinavir is a competitive inhibitor of the HIV protease by reversibly binding to the active site of the enzyme, preventing it from interacting with its substrate to produce mature and infectious viral particles. | |
| Record name | Nelfinavir | |
| Source | DrugBank | |
| URL | https://www.drugbank.ca/drugs/DB00220 | |
| Description | The DrugBank database is a unique bioinformatics and cheminformatics resource that combines detailed drug (i.e. chemical, pharmacological and pharmaceutical) data with comprehensive drug target (i.e. sequence, structure, and pathway) information. | |
| Explanation | Creative Common's Attribution-NonCommercial 4.0 International License (http://creativecommons.org/licenses/by-nc/4.0/legalcode) | |
CAS No. |
159989-64-7 | |
| Record name | Nelfinavir | |
| Source | CAS Common Chemistry | |
| URL | https://commonchemistry.cas.org/detail?cas_rn=159989-64-7 | |
| Description | CAS Common Chemistry is an open community resource for accessing chemical information. Nearly 500,000 chemical substances from CAS REGISTRY cover areas of community interest, including common and frequently regulated chemicals, and those relevant to high school and undergraduate chemistry classes. This chemical information, curated by our expert scientists, is provided in alignment with our mission as a division of the American Chemical Society. | |
| Explanation | The data from CAS Common Chemistry is provided under a CC-BY-NC 4.0 license, unless otherwise stated. | |
| Record name | Nelfinavir [INN:BAN] | |
| Source | ChemIDplus | |
| URL | https://pubchem.ncbi.nlm.nih.gov/substance/?source=chemidplus&sourceid=0159989647 | |
| Description | ChemIDplus is a free, web search system that provides access to the structure and nomenclature authority files used for the identification of chemical substances cited in National Library of Medicine (NLM) databases, including the TOXNET system. | |
| Record name | Nelfinavir | |
| Source | DrugBank | |
| URL | https://www.drugbank.ca/drugs/DB00220 | |
| Description | The DrugBank database is a unique bioinformatics and cheminformatics resource that combines detailed drug (i.e. chemical, pharmacological and pharmaceutical) data with comprehensive drug target (i.e. sequence, structure, and pathway) information. | |
| Explanation | Creative Common's Attribution-NonCommercial 4.0 International License (http://creativecommons.org/licenses/by-nc/4.0/legalcode) | |
| Record name | nelfinavir | |
| Source | DTP/NCI | |
| URL | https://dtp.cancer.gov/dtpstandard/servlet/dwindex?searchtype=NSC&outputformat=html&searchlist=747167 | |
| Description | The NCI Development Therapeutics Program (DTP) provides services and resources to the academic and private-sector research communities worldwide to facilitate the discovery and development of new cancer therapeutic agents. | |
| Explanation | Unless otherwise indicated, all text within NCI products is free of copyright and may be reused without our permission. Credit the National Cancer Institute as the source. | |
| Record name | Nelfinavir | |
| Source | EPA DSSTox | |
| URL | https://comptox.epa.gov/dashboard/DTXSID5035080 | |
| Description | DSSTox provides a high quality public chemistry resource for supporting improved predictive toxicology. | |
| Record name | NELFINAVIR | |
| Source | FDA Global Substance Registration System (GSRS) | |
| URL | https://gsrs.ncats.nih.gov/ginas/app/beta/substances/HO3OGH5D7I | |
| Description | The FDA Global Substance Registration System (GSRS) enables the efficient and accurate exchange of information on what substances are in regulated products. Instead of relying on names, which vary across regulatory domains, countries, and regions, the GSRS knowledge base makes it possible for substances to be defined by standardized, scientific descriptions. | |
| Explanation | Unless otherwise noted, the contents of the FDA website (www.fda.gov), both text and graphics, are not copyrighted. They are in the public domain and may be republished, reprinted and otherwise used freely by anyone without the need to obtain permission from FDA. Credit to the U.S. Food and Drug Administration as the source is appreciated but not required. | |
| Record name | Nelfinavir | |
| Source | Human Metabolome Database (HMDB) | |
| URL | http://www.hmdb.ca/metabolites/HMDB0014365 | |
| Description | The Human Metabolome Database (HMDB) is a freely available electronic database containing detailed information about small molecule metabolites found in the human body. | |
| Explanation | HMDB is offered to the public as a freely available resource. Use and re-distribution of the data, in whole or in part, for commercial purposes requires explicit permission of the authors and explicit acknowledgment of the source material (HMDB) and the original publication (see the HMDB citing page). We ask that users who download significant portions of the database cite the HMDB paper in any resulting publications. | |
Melting Point |
349.84 °C | |
| Record name | Nelfinavir | |
| Source | DrugBank | |
| URL | https://www.drugbank.ca/drugs/DB00220 | |
| Description | The DrugBank database is a unique bioinformatics and cheminformatics resource that combines detailed drug (i.e. chemical, pharmacological and pharmaceutical) data with comprehensive drug target (i.e. sequence, structure, and pathway) information. | |
| Explanation | Creative Common's Attribution-NonCommercial 4.0 International License (http://creativecommons.org/licenses/by-nc/4.0/legalcode) | |
| Record name | Nelfinavir | |
| Source | Human Metabolome Database (HMDB) | |
| URL | http://www.hmdb.ca/metabolites/HMDB0014365 | |
| Description | The Human Metabolome Database (HMDB) is a freely available electronic database containing detailed information about small molecule metabolites found in the human body. | |
| Explanation | HMDB is offered to the public as a freely available resource. Use and re-distribution of the data, in whole or in part, for commercial purposes requires explicit permission of the authors and explicit acknowledgment of the source material (HMDB) and the original publication (see the HMDB citing page). We ask that users who download significant portions of the database cite the HMDB paper in any resulting publications. | |
Retrosynthesis Analysis
AI-Powered Synthesis Planning: Our tool employs the Template_relevance Pistachio, Template_relevance Bkms_metabolic, Template_relevance Pistachio_ringbreaker, Template_relevance Reaxys, Template_relevance Reaxys_biocatalysis model, leveraging a vast database of chemical reactions to predict feasible synthetic routes.
One-Step Synthesis Focus: Specifically designed for one-step synthesis, it provides concise and direct routes for your target compounds, streamlining the synthesis process.
Accurate Predictions: Utilizing the extensive PISTACHIO, BKMS_METABOLIC, PISTACHIO_RINGBREAKER, REAXYS, REAXYS_BIOCATALYSIS database, our tool offers high-accuracy predictions, reflecting the latest in chemical research and data.
Strategy Settings
| Precursor scoring | Relevance Heuristic |
|---|---|
| Min. plausibility | 0.01 |
| Model | Template_relevance |
| Template Set | Pistachio/Bkms_metabolic/Pistachio_ringbreaker/Reaxys/Reaxys_biocatalysis |
| Top-N result to add to graph | 6 |
Feasible Synthetic Routes
Disclaimer and Information on In-Vitro Research Products
Please be aware that all articles and product information presented on BenchChem are intended solely for informational purposes. The products available for purchase on BenchChem are specifically designed for in-vitro studies, which are conducted outside of living organisms. In-vitro studies, derived from the Latin term "in glass," involve experiments performed in controlled laboratory settings using cells or tissues. It is important to note that these products are not categorized as medicines or drugs, and they have not received approval from the FDA for the prevention, treatment, or cure of any medical condition, ailment, or disease. We must emphasize that any form of bodily introduction of these products into humans or animals is strictly prohibited by law. It is essential to adhere to these guidelines to ensure compliance with legal and ethical standards in research and experimentation.