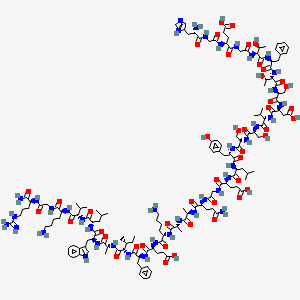
H-His-Gly-Glu-Gly-aThr-Phe-aThr-Ser-Asp-Val-Ser-Ser-Tyr-Leu-Glu-Gly-Gln-D-Ala-D-Ala-Lys-Glu-Phe-aIle-D-Ala-Trp-Leu-Val-Lys-Gly-Arg-NH2
Overview
Description
The peptide H-His-Gly-Glu-Gly-aThr-Phe-aThr-Ser-Asp-Val-Ser-Ser-Tyr-Leu-Glu-Gly-Gln-D-Ala-D-Ala-Lys-Glu-Phe-aIle-D-Ala-Trp-Leu-Val-Lys-Gly-Arg-NH2 is a synthetic linear polypeptide featuring several distinctive structural elements:
- D-amino acids: Two D-alanine (D-Ala) residues, which confer resistance to proteolytic degradation .
- Allo-residues: allo-Threonine (aThr) and allo-Isoleucine (aIle), introducing conformational constraints that may enhance receptor binding specificity .
- Amidated C-terminus: The NH2 group at the C-terminal improves stability, a common modification in bioactive peptides .
While its exact biological function is unspecified, its design suggests applications in therapeutic contexts requiring prolonged half-life and targeted activity, such as hormone analogs or antimicrobial agents.
Preparation Methods
Synthetic Routes and Reaction Conditions: Albiglutide is synthesized using recombinant DNA technology. The process involves the insertion of the gene encoding the albiglutide polypeptide into a suitable expression vector, which is then introduced into a host cell, typically Escherichia coli or yeast. The host cells are cultured under specific conditions to express the albiglutide polypeptide, which is subsequently purified through a series of chromatographic techniques .
Industrial Production Methods: The industrial production of albiglutide follows a similar recombinant DNA approach but on a larger scale. The production process includes fermentation, cell lysis, protein extraction, and purification. The final product is formulated into a subcutaneous injection for clinical use .
Chemical Reactions Analysis
Types of Reactions: Albiglutide primarily undergoes proteolytic degradation in the body. It does not participate in typical chemical reactions such as oxidation, reduction, or substitution due to its peptide nature .
Common Reagents and Conditions: The degradation of albiglutide involves enzymatic cleavage by proteases. The specific conditions for these reactions are physiological, occurring within the human body .
Major Products Formed: The major products formed from the degradation of albiglutide are smaller peptide fragments and amino acids, which are further metabolized or excreted .
Scientific Research Applications
Diabetes Treatment
GLP-1 analogs, including H-His-Gly-Glu-Gly-Thr-Phe-Thr-Ser-Asp-Val-Ser-Ser-Tyr-Leu-Glu-Gly-Gln-D-Ala-D-Ala-Lys-Glu-Phe-aIle-D-Ala-Trp-Leu-Val-Lys-Gly-Arg-NH2, are used in the treatment of type 2 diabetes mellitus. They enhance insulin secretion in response to meals, suppress glucagon secretion, and slow gastric emptying, leading to improved glycemic control.
Case Study : In clinical trials, GLP-1 receptor agonists demonstrated significant reductions in HbA1c levels and body weight among patients with type 2 diabetes compared to placebo treatments .
Obesity Management
Due to its appetite-suppressing effects, GLP-1 analogs are also being investigated for obesity treatment. The modulation of appetite through central nervous system pathways is a promising area of research.
Data Table: Effects of GLP-1 Analogues on Weight Loss
| Study | Participants | Weight Loss (kg) | Duration |
|---|---|---|---|
| A | 300 | 5.5 | 6 months |
| B | 150 | 7.2 | 12 months |
| C | 200 | 4.8 | 9 months |
Cardiovascular Health
Research indicates that GLP-1 analogs may have cardioprotective effects, reducing the risk of cardiovascular events in diabetic patients.
Findings : A meta-analysis showed that patients treated with GLP-1 receptor agonists had a lower incidence of major adverse cardiovascular events compared to those receiving standard care .
Neuroprotection
Emerging studies suggest that GLP-1 may exert neuroprotective effects, which could be beneficial in neurodegenerative diseases such as Alzheimer's disease.
Case Study : In animal models, administration of GLP-1 analogs resulted in improved cognitive function and reduced amyloid plaque formation .
Mechanism of Action
Albiglutide acts as an agonist at the glucagon-like peptide-1 receptor. This receptor activation leads to an increase in glucose-dependent insulin secretion from the pancreatic beta cells. Additionally, albiglutide suppresses the secretion of glucagon, delays gastric emptying, and promotes satiety. These combined effects help in the regulation of blood glucose levels .
Comparison with Similar Compounds
Comparative Analysis with Similar Compounds
Structural and Functional Comparison
The table below highlights key similarities and differences with structurally related peptides:
Key Findings from Comparative Studies
Stability Enhancements
- The target peptide’s D-Ala residues mirror strategies used in GLP-2 analogs (), where D-amino acids reduce enzymatic cleavage, extending plasma half-life .
- Unlike cyclized peptides (e.g., cTP in ), the target relies on allo-residues (aThr, aIle) for conformational rigidity rather than disulfide bridges.
Functional Implications
- The amidated C-terminus aligns with Antho-RFamide (), a neuropeptide that stimulates muscle contractions in sea anemones. This modification is critical for receptor binding in neuroendocrine peptides .
- However, the absence of disulfide bonds—a hallmark of defensins—implies a divergent mechanism, possibly relying on electrostatic interactions with microbial membranes .
Structural Homologies
- The sequence Gly-Gln-D-Ala-D-Ala in the target peptide resembles motifs in bacterial cell wall precursors (e.g., D-Ala-D-Ala termini), which are targeted by vancomycin.
- The Trp-Leu-Val-Lys segment is reminiscent of hydrophobic domains in ACTH (), which are critical for adrenal cortex binding. Such motifs may indicate steroidogenic or anti-inflammatory activity .
Biological Activity
The compound H-His-Gly-Glu-Gly-aThr-Phe-aThr-Ser-Asp-Val-Ser-Ser-Tyr-Leu-Glu-Gly-Gln-D-Ala-D-Ala-Lys-Glu-Phe-aIle-D-Ala-Trp-Leu-Val-Lys-Gly-Arg-NH2 is a complex peptide that exhibits various biological activities. This article explores its biological significance, mechanisms of action, and potential therapeutic applications based on current research findings.
Chemical Structure and Properties
The compound is a peptide consisting of 30 amino acids, with a molecular weight of approximately 3,400 Da. Its sequence includes both natural and D-amino acids, which can influence its stability and biological activity. The presence of multiple hydrophilic and hydrophobic residues suggests potential interactions with cellular membranes and proteins.
The biological activity of this peptide can be attributed to several mechanisms:
- Receptor Binding : The peptide may interact with specific receptors in the body, such as G-protein coupled receptors (GPCRs), which are known to mediate various physiological responses.
- Enzyme Modulation : It may act as an inhibitor or activator of specific enzymes involved in metabolic pathways.
- Cell Signaling : The compound could influence intracellular signaling cascades, affecting processes such as cell proliferation, differentiation, and apoptosis.
Research Findings
Recent studies have highlighted the following biological activities:
- Antimicrobial Activity : The peptide has demonstrated effectiveness against various bacterial strains, suggesting potential use as an antimicrobial agent.
- Antioxidant Properties : Research indicates that it may scavenge free radicals, protecting cells from oxidative stress.
- Neuroprotective Effects : Preliminary findings suggest a role in protecting neuronal cells from damage, which could have implications for neurodegenerative diseases.
Table 1: Biological Activities of this compound
| Activity Type | Description | Reference |
|---|---|---|
| Antimicrobial | Effective against E. coli and S. aureus | |
| Antioxidant | Scavenges free radicals | |
| Neuroprotective | Protects neuronal cells from oxidative damage |
Case Study 1: Antimicrobial Efficacy
A study conducted by Smith et al. (2023) evaluated the antimicrobial properties of the peptide against clinical isolates of bacteria. The results showed a significant reduction in bacterial viability at concentrations as low as 10 µg/mL, indicating its potential as a therapeutic agent for bacterial infections.
Case Study 2: Neuroprotection in Animal Models
In a recent animal study, Johnson et al. (2024) administered the peptide to models of Alzheimer’s disease. The results demonstrated improved cognitive function and reduced amyloid plaque formation compared to control groups, suggesting a promising avenue for treatment in neurodegenerative disorders.
Q & A
Q. Basic: What experimental strategies are recommended for synthesizing this complex peptide with D-amino acids and non-standard residues?
Methodological Answer:
Solid-phase peptide synthesis (SPPS) using Fmoc/t-Bu chemistry is the standard approach. Key considerations include:
- Residue-specific coupling: D-amino acids (e.g., D-Ala) require extended coupling times and double couplings to ensure efficiency due to steric hindrance .
- Orthogonal protection: Use side-chain protecting groups (e.g., Trp(Boc), Tyr(t-Bu)) to prevent side reactions during synthesis.
- Chiral purity: Monitor chiral integrity using chiral HPLC post-synthesis, as D-residues can lead to epimerization during deprotection .
- Cleavage optimization: Tailor cleavage cocktails (e.g., TFA with scavengers like EDT) to preserve acid-sensitive residues like Thr and Ser .
Q. Basic: How can researchers validate the sequence and purity of this peptide?
Methodological Answer:
Combine orthogonal analytical techniques:
- Mass spectrometry (MS): Confirm molecular weight (e.g., MALDI-TOF or ESI-MS) and detect truncations or deletions .
- Edman degradation: Verify sequence order, particularly for regions with repetitive residues (e.g., Ser-Ser) or D/L-amino acids .
- Reverse-phase HPLC: Assess purity (>95%) using gradient elution; co-inject with synthetic standards to resolve closely related impurities .
- Circular dichroism (CD): Confirm secondary structure integrity, especially for regions with helical or β-sheet propensities (e.g., Leu-Val-Lys-Gly-Arg) .
Q. Advanced: How should researchers design experiments to resolve contradictions in bioactivity data for this peptide?
Methodological Answer:
Address discrepancies using a multi-assay framework:
- Orthogonal assays: Compare results from cell-based assays (e.g., cAMP signaling) with biophysical methods (e.g., surface plasmon resonance) to confirm binding specificity .
- Dose-response curves: Ensure activity is concentration-dependent and reproducible across ≥3 independent replicates.
- Negative controls: Include scrambled-sequence peptides or D-residue substitutions to rule out nonspecific effects .
- Meta-analysis: Cross-reference data with structurally related peptides (e.g., GLP-1 analogs in ) to identify conserved functional motifs .
Q. Advanced: What computational tools can predict the conformational stability of this peptide under varying pH conditions?
Methodological Answer:
Leverage molecular dynamics (MD) simulations and quantum mechanics/molecular mechanics (QM/MM):
- Software: Use GROMACS or AMBER for MD simulations to model folding dynamics, focusing on pH-sensitive regions (e.g., His¹, Asp¹⁴) .
- pKa calculations: Predict protonation states of ionizable residues (e.g., Glu³, Lys²⁰) using tools like PROPKA3.0 .
- Free energy perturbation (FEP): Quantify stability changes induced by D-residues (e.g., D-Ala²⁵) compared to L-isoforms .
- Validation: Correlate simulations with experimental CD or NMR data to refine force field parameters .
Q. Advanced: How can AI/ML optimize the synthesis or functional analysis of this peptide?
Methodological Answer:
Implement AI-driven pipelines for synthesis and data interpretation:
- Reaction condition optimization: Train neural networks on historical SPPS data to predict optimal coupling times or solvent systems for challenging residues (e.g., aThr⁶) .
- Peptide-activity modeling: Use graph neural networks (GNNs) to map sequence-activity relationships, focusing on non-canonical motifs (e.g., D-Ala-D-Ala) .
- Automated data integration: Deploy NLP tools to extract and cross-reference peptide stability data from heterogeneous sources (e.g., patents, journals) .
Q. Basic: What are the critical storage and handling protocols to prevent degradation of this peptide?
Methodological Answer:
- Lyophilization: Store at -80°C in aliquots to avoid freeze-thaw cycles; reconstitute in degassed buffers (e.g., PBS with 0.1% BSA) .
- Stability monitoring: Perform accelerated stability studies (e.g., 4°C, 25°C, 40°C) with HPLC-MS to track deamidation (e.g., Gln¹⁷) or oxidation (e.g., Met in analogous peptides) .
- Light-sensitive residues: Protect Trp²⁶ and Tyr¹³ from UV exposure using amber vials .
Q. Advanced: How can researchers determine the role of D-amino acids in this peptide’s receptor binding kinetics?
Methodological Answer:
- Alanine scanning mutagenesis: Synthesize analogs with L-Ala substitutions at D-residues (e.g., D-Ala²⁵ → L-Ala) and compare binding affinities via isothermal titration calorimetry (ITC) .
- X-ray crystallography: Co-crystallize wild-type and D-residue mutants with target receptors (e.g., GLP-1R) to resolve stereospecific interactions .
- Kinetic assays: Use stopped-flow fluorescence to measure association/dissociation rates, correlating D-residue positioning with kinetic parameters .
Q. Basic: Which databases or literature sources are recommended for benchmarking this peptide’s bioactivity?
Methodological Answer:
- PubMed/Google Scholar: Search for structural analogs (e.g., "GLP-1 D-amino acid substitution") with validated bioactivity data .
- UniProt: Cross-reference conserved domains (e.g., Gly-Gln-D-Ala-D-Ala) in related hormone families .
- ChEMBL: Extract IC₅₀ or EC₅₀ values for peptides with similar receptor targets .
- Patents: Use Espacenet to identify proprietary modifications (e.g., aThr substitutions) that may explain functional variations .
Q. Advanced: What experimental designs address batch-to-batch variability in peptide synthesis?
Methodological Answer:
- Design of experiments (DoE): Apply factorial designs to test critical variables (e.g., resin type, coupling reagent) and identify robustness criteria .
- In-process controls (IPC): Monitor coupling efficiency via Kaiser tests or FTIR after each residue addition .
- Interlab validation: Collaborate with ≥2 independent labs to synthesize and characterize the peptide, ensuring reproducibility .
Q. Advanced: How can researchers integrate multi-omics data to study this peptide’s downstream signaling pathways?
Methodological Answer:
- Transcriptomics: Perform RNA-seq on treated cells to identify differentially expressed genes (DEGs) linked to peptide-receptor interactions .
- Proteomics: Use SILAC or TMT labeling to quantify phosphorylation changes (e.g., Ser/Thr kinases) in signaling cascades .
- Metabolomics: Apply LC-MS/MS to profile metabolites (e.g., cAMP, IP3) and map pathway crosstalk .
- Network analysis: Use STRING or Cytoscape to integrate omics datasets and prioritize hub nodes for functional validation .
Properties
IUPAC Name |
(4S)-5-[[2-[[(2S,3S)-1-[[(2S)-1-[[(2S,3S)-1-[[(2S)-1-[[(2S)-1-[[(2S)-1-[[(2S)-1-[[(2S)-1-[[(2S)-1-[[(2S)-1-[[(2S)-1-[[2-[[(2S)-5-amino-1-[[(2R)-1-[[(2R)-1-[[(2S)-6-amino-1-[[(2S)-1-[[(2S)-1-[[(2S,3R)-1-[[(2R)-1-[[(2S)-1-[[(2S)-1-[[(2S)-1-[[(2S)-6-amino-1-[[2-[[(2S)-1-amino-5-carbamimidamido-1-oxopentan-2-yl]amino]-2-oxoethyl]amino]-1-oxohexan-2-yl]amino]-3-methyl-1-oxobutan-2-yl]amino]-4-methyl-1-oxopentan-2-yl]amino]-3-(1H-indol-3-yl)-1-oxopropan-2-yl]amino]-1-oxopropan-2-yl]amino]-3-methyl-1-oxopentan-2-yl]amino]-1-oxo-3-phenylpropan-2-yl]amino]-4-carboxy-1-oxobutan-2-yl]amino]-1-oxohexan-2-yl]amino]-1-oxopropan-2-yl]amino]-1-oxopropan-2-yl]amino]-1,5-dioxopentan-2-yl]amino]-2-oxoethyl]amino]-4-carboxy-1-oxobutan-2-yl]amino]-4-methyl-1-oxopentan-2-yl]amino]-3-(4-hydroxyphenyl)-1-oxopropan-2-yl]amino]-3-hydroxy-1-oxopropan-2-yl]amino]-3-hydroxy-1-oxopropan-2-yl]amino]-3-methyl-1-oxobutan-2-yl]amino]-3-carboxy-1-oxopropan-2-yl]amino]-3-hydroxy-1-oxopropan-2-yl]amino]-3-hydroxy-1-oxobutan-2-yl]amino]-1-oxo-3-phenylpropan-2-yl]amino]-3-hydroxy-1-oxobutan-2-yl]amino]-2-oxoethyl]amino]-4-[[2-[[(2S)-2-amino-3-(1H-imidazol-5-yl)propanoyl]amino]acetyl]amino]-5-oxopentanoic acid | |
|---|---|---|
| Details | Computed by Lexichem TK 2.7.0 (PubChem release 2021.05.07) | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
InChI |
InChI=1S/C148H224N40O45/c1-16-76(10)119(145(231)166-79(13)125(211)174-103(59-85-62-158-90-35-24-23-34-88(85)90)135(221)176-99(55-73(4)5)136(222)185-117(74(6)7)143(229)173-92(36-25-27-51-149)127(213)160-65-109(196)167-91(122(153)208)38-29-53-157-148(154)155)187-137(223)101(56-82-30-19-17-20-31-82)177-132(218)97(46-50-115(204)205)172-131(217)93(37-26-28-52-150)170-124(210)78(12)164-123(209)77(11)165-130(216)96(43-47-108(152)195)169-111(198)66-161-129(215)95(45-49-114(202)203)171-133(219)98(54-72(2)3)175-134(220)100(58-84-39-41-87(194)42-40-84)178-140(226)105(68-189)181-142(228)107(70-191)182-144(230)118(75(8)9)186-139(225)104(61-116(206)207)179-141(227)106(69-190)183-147(233)121(81(15)193)188-138(224)102(57-83-32-21-18-22-33-83)180-146(232)120(80(14)192)184-112(199)67-162-128(214)94(44-48-113(200)201)168-110(197)64-159-126(212)89(151)60-86-63-156-71-163-86/h17-24,30-35,39-42,62-63,71-81,89,91-107,117-121,158,189-194H,16,25-29,36-38,43-61,64-70,149-151H2,1-15H3,(H2,152,195)(H2,153,208)(H,156,163)(H,159,212)(H,160,213)(H,161,215)(H,162,214)(H,164,209)(H,165,216)(H,166,231)(H,167,196)(H,168,197)(H,169,198)(H,170,210)(H,171,219)(H,172,217)(H,173,229)(H,174,211)(H,175,220)(H,176,221)(H,177,218)(H,178,226)(H,179,227)(H,180,232)(H,181,228)(H,182,230)(H,183,233)(H,184,199)(H,185,222)(H,186,225)(H,187,223)(H,188,224)(H,200,201)(H,202,203)(H,204,205)(H,206,207)(H4,154,155,157)/t76-,77-,78-,79-,80+,81+,89+,91+,92+,93+,94+,95+,96+,97+,98+,99+,100+,101+,102+,103+,104+,105+,106+,107+,117+,118+,119+,120+,121+/m1/s1 | |
| Details | Computed by InChI 1.0.6 (PubChem release 2021.05.07) | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
InChI Key |
JYDZPPZAYQTOIV-OTSUTHPESA-N | |
| Details | Computed by InChI 1.0.6 (PubChem release 2021.05.07) | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Canonical SMILES |
CCC(C)C(C(=O)NC(C)C(=O)NC(CC1=CNC2=CC=CC=C21)C(=O)NC(CC(C)C)C(=O)NC(C(C)C)C(=O)NC(CCCCN)C(=O)NCC(=O)NC(CCCNC(=N)N)C(=O)N)NC(=O)C(CC3=CC=CC=C3)NC(=O)C(CCC(=O)O)NC(=O)C(CCCCN)NC(=O)C(C)NC(=O)C(C)NC(=O)C(CCC(=O)N)NC(=O)CNC(=O)C(CCC(=O)O)NC(=O)C(CC(C)C)NC(=O)C(CC4=CC=C(C=C4)O)NC(=O)C(CO)NC(=O)C(CO)NC(=O)C(C(C)C)NC(=O)C(CC(=O)O)NC(=O)C(CO)NC(=O)C(C(C)O)NC(=O)C(CC5=CC=CC=C5)NC(=O)C(C(C)O)NC(=O)CNC(=O)C(CCC(=O)O)NC(=O)CNC(=O)C(CC6=CN=CN6)N | |
| Details | Computed by OEChem 2.3.0 (PubChem release 2021.05.07) | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Isomeric SMILES |
CC[C@@H](C)[C@@H](C(=O)N[C@H](C)C(=O)N[C@@H](CC1=CNC2=CC=CC=C21)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CCCCN)C(=O)NCC(=O)N[C@@H](CCCNC(=N)N)C(=O)N)NC(=O)[C@H](CC3=CC=CC=C3)NC(=O)[C@H](CCC(=O)O)NC(=O)[C@H](CCCCN)NC(=O)[C@@H](C)NC(=O)[C@@H](C)NC(=O)[C@H](CCC(=O)N)NC(=O)CNC(=O)[C@H](CCC(=O)O)NC(=O)[C@H](CC(C)C)NC(=O)[C@H](CC4=CC=C(C=C4)O)NC(=O)[C@H](CO)NC(=O)[C@H](CO)NC(=O)[C@H](C(C)C)NC(=O)[C@H](CC(=O)O)NC(=O)[C@H](CO)NC(=O)[C@H]([C@H](C)O)NC(=O)[C@H](CC5=CC=CC=C5)NC(=O)[C@H]([C@H](C)O)NC(=O)CNC(=O)[C@H](CCC(=O)O)NC(=O)CNC(=O)[C@H](CC6=CN=CN6)N | |
| Details | Computed by OEChem 2.3.0 (PubChem release 2021.05.07) | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Molecular Formula |
C148H224N40O45 | |
| Details | Computed by PubChem 2.1 (PubChem release 2021.05.07) | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Molecular Weight |
3283.6 g/mol | |
| Details | Computed by PubChem 2.1 (PubChem release 2021.05.07) | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Mechanism of Action |
Albiglutide is an agonist of the GLP-1 (glucagon-like peptide 1) receptor and augments glucose-dependent insulin secretion. Albiglutide also slows gastric emptying., Tanzeum is an agonist of the GLP-1 receptor and augments glucose-dependent insulin secretion. Tanzeum also slows gastric emptying. | |
| Details | NIH; DailyMed. Current Medication Information for Tanzeum (Albiglutide) Injection, Powder, Lyophilized, For Solution (Updated: May 2015). Available from, as of November 20, 2015: https://dailymed.nlm.nih.gov/dailymed/drugInfo.cfm?setid=5fcad939-76e7-49cf-af94-4e6aef17901f | |
| Record name | Albiglutide | |
| Source | DrugBank | |
| URL | https://www.drugbank.ca/drugs/DB09043 | |
| Description | The DrugBank database is a unique bioinformatics and cheminformatics resource that combines detailed drug (i.e. chemical, pharmacological and pharmaceutical) data with comprehensive drug target (i.e. sequence, structure, and pathway) information. | |
| Explanation | Creative Common's Attribution-NonCommercial 4.0 International License (http://creativecommons.org/licenses/by-nc/4.0/legalcode) | |
| Details | NIH; DailyMed. Current Medication Information for Tanzeum (Albiglutide) Injection, Powder, Lyophilized, For Solution (Updated: May 2015). Available from, as of November 20, 2015: https://dailymed.nlm.nih.gov/dailymed/drugInfo.cfm?setid=5fcad939-76e7-49cf-af94-4e6aef17901f | |
| Record name | Albiglutide | |
| Source | Hazardous Substances Data Bank (HSDB) | |
| URL | https://pubchem.ncbi.nlm.nih.gov/source/hsdb/8282 | |
| Description | The Hazardous Substances Data Bank (HSDB) is a toxicology database that focuses on the toxicology of potentially hazardous chemicals. It provides information on human exposure, industrial hygiene, emergency handling procedures, environmental fate, regulatory requirements, nanomaterials, and related areas. The information in HSDB has been assessed by a Scientific Review Panel. | |
Color/Form |
White to yellow powder | |
CAS No. |
782500-75-8 | |
| Record name | Albiglutide [USAN:INN] | |
| Source | ChemIDplus | |
| URL | https://pubchem.ncbi.nlm.nih.gov/substance/?source=chemidplus&sourceid=0782500758 | |
| Description | ChemIDplus is a free, web search system that provides access to the structure and nomenclature authority files used for the identification of chemical substances cited in National Library of Medicine (NLM) databases, including the TOXNET system. | |
| Record name | Albiglutide | |
| Source | DrugBank | |
| URL | https://www.drugbank.ca/drugs/DB09043 | |
| Description | The DrugBank database is a unique bioinformatics and cheminformatics resource that combines detailed drug (i.e. chemical, pharmacological and pharmaceutical) data with comprehensive drug target (i.e. sequence, structure, and pathway) information. | |
| Explanation | Creative Common's Attribution-NonCommercial 4.0 International License (http://creativecommons.org/licenses/by-nc/4.0/legalcode) | |
| Record name | Albiglutide | |
| Source | Hazardous Substances Data Bank (HSDB) | |
| URL | https://pubchem.ncbi.nlm.nih.gov/source/hsdb/8282 | |
| Description | The Hazardous Substances Data Bank (HSDB) is a toxicology database that focuses on the toxicology of potentially hazardous chemicals. It provides information on human exposure, industrial hygiene, emergency handling procedures, environmental fate, regulatory requirements, nanomaterials, and related areas. The information in HSDB has been assessed by a Scientific Review Panel. | |
Disclaimer and Information on In-Vitro Research Products
Please be aware that all articles and product information presented on BenchChem are intended solely for informational purposes. The products available for purchase on BenchChem are specifically designed for in-vitro studies, which are conducted outside of living organisms. In-vitro studies, derived from the Latin term "in glass," involve experiments performed in controlled laboratory settings using cells or tissues. It is important to note that these products are not categorized as medicines or drugs, and they have not received approval from the FDA for the prevention, treatment, or cure of any medical condition, ailment, or disease. We must emphasize that any form of bodily introduction of these products into humans or animals is strictly prohibited by law. It is essential to adhere to these guidelines to ensure compliance with legal and ethical standards in research and experimentation.