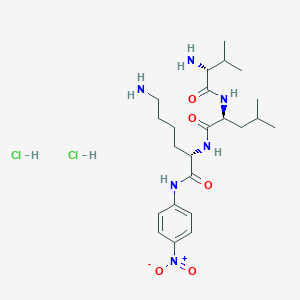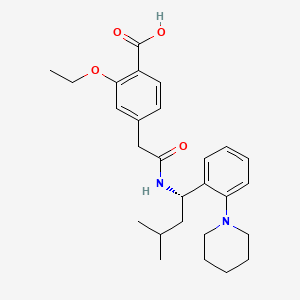
Repaglinide
概述
准备方法
合成路线和反应条件
瑞格列奈可以通过涉及多个关键中间体的多步合成过程来合成。 一种常见的合成方法包括以下步骤 :
酯化: 3-羟基苯乙酸被酯化生成3-羟基苯乙酸乙酯。
甲酰化: 然后将酯甲酰化以生成3-甲酰基-4-羟基苯乙酸乙酯。
氧化: 甲酰基被氧化成羧酸,生成3-羧基-4-羟基苯乙酸乙酯。
醚化: 羟基被醚化生成3-羧基-4-乙氧基苯乙酸乙酯。
选择性水解: 选择性水解酯,生成3-羧基-4-乙氧基苯乙酸,它是瑞格列奈合成的关键中间体。
工业生产方法
瑞格列奈的工业生产涉及优化反应条件以最大化收率和纯度。 这包括控制反应温度、时间、溶剂和底物比率 . 该过程旨在可扩展且环境友好,杂质最少。
化学反应分析
反应类型
瑞格列奈会经历多种类型的化学反应,包括:
氧化: 瑞格列奈可以被氧化形成各种代谢产物。
还原: 还原反应可以改变瑞格列奈分子上的官能团。
取代: 取代反应可以在芳香环和侧链的不同位置发生。
常见试剂和条件
用于合成和修饰瑞格列奈的常见试剂包括:
氧化剂: 例如高锰酸钾和过氧化氢。
还原剂: 例如硼氢化钠和氢化锂铝。
取代试剂: 例如卤素和烷基化剂。
主要产物
从这些反应中形成的主要产物包括瑞格列奈的各种代谢产物和衍生物,可以使用 FT-IR、NMR 和 UV-Vis 光谱等技术对其进行表征 .
科学研究应用
Repaglinide is an oral antihyperglycemic agent primarily used to manage type 2 diabetes mellitus . It helps regulate blood glucose levels by stimulating the pancreas to release insulin . It is particularly effective in controlling postprandial glucose levels .
Monotherapy
This compound can be used as a monotherapy to manage blood sugar levels. A study comparing this compound monotherapy to nateglinide monotherapy showed that this compound was more effective in reducing HbA1c and fasting plasma glucose (FPG) values after 16 weeks of treatment . After 16 weeks, the mean reduction in HbA1c was significantly greater with this compound than with nateglinide (-1.57% vs -1.04%; P=0.002). Additionally, the mean changes in FPG demonstrated greater efficacy for this compound than nateglinide (-57 vs -18 mg/dl; P<0.001) .
Combination Therapy
This compound can be used in combination with other oral anti-diabetic drugs . this compound is also effective in lowering blood glucose concentrations in both sulfonylurea-treated patients and those who are new to oral hypoglycemic agents .
Glycemic Efficacy
Studies have demonstrated that reductions in A1c levels achieved with meglitinides like this compound are similar to, or slightly less than, those observed with sulfonylurea or metformin treatment when used as monotherapy . In a study comparing this compound to nateglinide in patients with type 2 diabetes, this compound was more effective in reducing A1c levels (1.57% vs. 1.04%) .
Safety and Tolerability
This compound is generally well-tolerated . However, like sulfonylureas, meglitinides can cause hypoglycemia, although the risk of severe hypoglycemia is less . Weight gain is another potential side effect .
Dosage and Administration
The recommended starting dose of this compound for patients with an A1c less than 8% is 0.5 mg before each meal (1-30 minutes prior). For those with an A1c of 8% or greater, the starting dose is 1 or 2 mg orally before each meal. The dose can be doubled up to 4mg with each meal until satisfactory glycemic control is achieved, with a one-week interval between dose increases. The maximum daily dose is 16 mg per day . this compound should be taken with meals, and doses should be skipped if a meal is skipped .
This compound vs. Nateglinide
作用机制
瑞格列奈通过与胰腺β细胞上的 ATP 敏感性钾通道结合发挥作用。这种结合抑制钾离子的外流,导致细胞膜去极化。去极化打开电压依赖性钙通道,使钙离子进入细胞。 钙离子的流入触发胰岛素颗粒的外排,导致胰岛素释放增加 . 这种机制依赖于葡萄糖,这意味着只有在存在葡萄糖的情况下才会刺激胰岛素释放,从而降低低血糖的风险 .
相似化合物的比较
瑞格列奈经常与其他抗糖尿病药物进行比较,例如:
那格列奈: 另一种甲磺酰脲类药物,具有类似的作用机制,但作用持续时间更短。
磺酰脲类药物: 例如格列本脲和格列美脲,它们也会刺激胰岛素释放,但作用持续时间更长,发生低血糖的风险更高。
生物活性
Repaglinide is an oral antidiabetic medication belonging to the class of meglitinides, primarily used for managing type 2 diabetes mellitus. Its mechanism of action involves stimulating insulin secretion from pancreatic β-cells in response to elevated blood glucose levels. This article delves into the biological activity of this compound, supported by data tables, case studies, and detailed research findings.
This compound functions by inhibiting ATP-sensitive potassium channels (K-ATP channels) in pancreatic β-cells. This inhibition leads to membrane depolarization, opening L-type calcium channels, and subsequently increasing intracellular calcium concentrations. The rise in calcium levels triggers the exocytosis of insulin granules, enhancing insulin secretion in a glucose-dependent manner. Unlike sulfonylureas, this compound's effect is minimal in the absence of glucose, making it particularly effective for postprandial blood glucose control .
Pharmacokinetics
The pharmacokinetic profile of this compound is characterized by rapid absorption and a short duration of action:
- Absorption : Rapidly absorbed with peak plasma concentrations reached within 0.5 to 1.4 hours after oral administration.
- Bioavailability : Approximately 56%, indicating that a significant portion of the drug reaches systemic circulation.
- Volume of Distribution : About 31 L following intravenous administration.
- Protein Binding : Over 98%, primarily to albumin and α1-acid glycoprotein .
Case Studies and Clinical Trials
-
Efficacy in Type 2 Diabetes Management :
A phase II multicenter trial involving 99 patients demonstrated that this compound significantly reduced glycosylated hemoglobin (HbA1c) levels from 8.5% to 7.8% compared to an increase from 8.1% to 9.3% in the placebo group (P < 0.0001). Additionally, fasting plasma glucose levels decreased significantly in the this compound group . -
Comparison with Other Antidiabetics :
In a study comparing this compound with nateglinide over 16 weeks, this compound resulted in a greater reduction in HbA1c (1.57% vs. 1.04%, P = 0.002) and was more effective at controlling postprandial glucose levels . -
Pharmacogenomic Studies :
Research involving newly diagnosed type 2 diabetes patients indicated that genetic factors could influence the efficacy of this compound, with significant associations found between genetic scores and reductions in fasting glucose and HbA1c levels .
Safety Profile
This compound is generally well tolerated, with common adverse effects including:
- Hypoglycemia : Occurred in approximately 16% of patients, similar to other sulfonylureas but without increased risk when meals were missed.
- Other Adverse Events : Upper respiratory infections (10%), rhinitis (7%), bronchitis (6%), and headache (9%) were also reported .
Summary Table of Key Findings
| Parameter | Value/Outcome |
|---|---|
| Mechanism of Action | Inhibition of K-ATP channels leading to insulin secretion |
| Peak Plasma Concentration | 0.5 - 1.4 hours after administration |
| Bioavailability | ~56% |
| Reduction in HbA1c | From 8.5% to 7.8% (P < 0.0001) |
| Common Adverse Effects | Hypoglycemia (16%), upper respiratory infection (10%) |
常见问题
Basic Research Questions
Q. What are the key methodological considerations when designing preclinical studies to evaluate repaglinide’s pharmacokinetics and safety?
- Methodological Answer : Preclinical studies must adhere to NIH guidelines for experimental rigor and reproducibility. Key factors include:
- Sample Size and Power Analysis : Use statistical tools (e.g., G*Power) to determine adequate sample sizes, minimizing Type I/II errors .
- In Vivo Model Selection : Rodent models should be justified based on metabolic similarities to humans, with documentation of strain, age, and housing conditions .
- Dosage Rationale : Align with human-equivalent doses (HED) derived from body surface area scaling, as per FDA guidance.
- Ethical Compliance : Include Institutional Animal Care and Use Committee (IACUC) approval details .
Q. How can researchers formulate focused research questions for optimizing this compound formulations using frameworks like PICO or FINER?
- Methodological Answer :
- PICO Framework : Define Population (e.g., diabetic rat models), Intervention (e.g., floating tablets with HPMC K-100M), Comparison (e.g., immediate-release formulations), and Outcome (e.g., gastric retention time) .
- FINER Criteria : Ensure questions are Feasible (e.g., lab resources for sustained-release testing), Interesting (e.g., novel polymer combinations), Novel (e.g., addressing gaps in floating mechanisms), Ethical, and Relevant (e.g., clinical translation potential) .
Q. What strategies ensure rigorous literature reviews for identifying gaps in this compound delivery systems?
- Methodological Answer :
- Systematic Search : Use databases (PubMed, Scopus) with Boolean operators (e.g., "this compound AND floating tablets NOT metformin").
- Quality Assessment : Apply PRISMA guidelines to filter studies by robustness (e.g., peer-reviewed journals, sample size >30) .
- Gap Analysis : Map existing studies using tools like VOSviewer to visualize under-researched areas (e.g., long-term stability of gastroretentive formulations) .
Advanced Research Questions
Q. How do response surface methodologies (RSM) such as Central Composite Design (CCD) and D-Optimal Design compare in optimizing this compound formulations?
- Methodological Answer :
- Model Accuracy : CCD typically achieves R² > 0.85 but requires more runs (e.g., 20 runs for 3 factors), whereas D-Optimal reduces runs (e.g., 15) with comparable predictive power (relative error < 4%) .
- Diagnostic Tools : Use ANOVA for model significance (p < 0.05), residual plots for variance homogeneity, and leverage 3D contour plots to visualize interactions (e.g., HPMC K-100M concentration vs. floating lag time) .
- Software : Implement Design-Expert® or JMP® for automated optimization, prioritizing "desirability functions" to balance conflicting responses (e.g., drug release vs. buoyancy) .
Q. What statistical approaches resolve contradictions in this compound’s pharmacokinetic data across heterogeneous patient cohorts?
- Methodological Answer :
- Mixed-Effects Modeling : Account for inter-individual variability (e.g., CYP2C8 polymorphism) using NONMEM® or Monolix® .
- Sensitivity Analysis : Identify covariates (e.g., BMI, renal function) via stepwise covariate modeling (SCM) .
- Meta-Regression : Pool data from multiple studies (e.g., Cochrane Library) to adjust for confounders like dosing regimens .
Q. How should researchers structure the Methods section to meet reproducibility standards for complex this compound formulations?
- Methodological Answer :
- Granularity : Specify excipient sources (e.g., Eudragit RSPO from Evonik Industries), tablet hardness (e.g., 8–10 kp), and dissolution conditions (e.g., USP Apparatus II, 50 rpm, 37°C) .
- Validation : Include HPLC parameters (column: C18, 5 µm; mobile phase: acetonitrile-phosphate buffer) and validation metrics (e.g., LOD/LOQ, recovery rate 98–102%) .
- Ethnography for Adherence Studies : For patient-centric research, detail interview protocols (e.g., semi-structured questionnaires) and coding frameworks (e.g., NVivo® for thematic analysis) .
Q. Data Analysis & Interpretation
Q. What advanced techniques validate the robustness of this compound’s in vitro-in vivo correlation (IVIVC)?
- Methodological Answer :
- Level A IVIVC : Use deconvolution methods (e.g., Wagner-Nelson) to correlate dissolution profiles (e.g., f2 similarity factor >50) with plasma concentration-time curves .
- Bootstrap Validation : Resample data 1000x to compute 95% confidence intervals for predicted AUC and Cmax .
Q. How can machine learning improve this compound formulation development?
- Methodological Answer :
- Feature Engineering : Input variables include polymer viscosity, drug-polymer ratio, and compression force.
- Algorithm Selection : Compare Random Forest (interpretability) vs. Neural Networks (nonlinearity handling) using metrics like RMSE .
- Validation : Apply k-fold cross-validation (k=10) to prevent overfitting .
属性
IUPAC Name |
2-ethoxy-4-[2-[[(1S)-3-methyl-1-(2-piperidin-1-ylphenyl)butyl]amino]-2-oxoethyl]benzoic acid | |
|---|---|---|
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
InChI |
InChI=1S/C27H36N2O4/c1-4-33-25-17-20(12-13-22(25)27(31)32)18-26(30)28-23(16-19(2)3)21-10-6-7-11-24(21)29-14-8-5-9-15-29/h6-7,10-13,17,19,23H,4-5,8-9,14-16,18H2,1-3H3,(H,28,30)(H,31,32)/t23-/m0/s1 | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
InChI Key |
FAEKWTJYAYMJKF-QHCPKHFHSA-N | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Canonical SMILES |
CCOC1=C(C=CC(=C1)CC(=O)NC(CC(C)C)C2=CC=CC=C2N3CCCCC3)C(=O)O | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Isomeric SMILES |
CCOC1=C(C=CC(=C1)CC(=O)N[C@@H](CC(C)C)C2=CC=CC=C2N3CCCCC3)C(=O)O | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Molecular Formula |
C27H36N2O4 | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
DSSTOX Substance ID |
DTXSID3023552 | |
| Record name | Repaglinide | |
| Source | EPA DSSTox | |
| URL | https://comptox.epa.gov/dashboard/DTXSID3023552 | |
| Description | DSSTox provides a high quality public chemistry resource for supporting improved predictive toxicology. | |
Molecular Weight |
452.6 g/mol | |
| Source | PubChem | |
| URL | https://pubchem.ncbi.nlm.nih.gov | |
| Description | Data deposited in or computed by PubChem | |
Physical Description |
Solid | |
| Record name | Repaglinide | |
| Source | Human Metabolome Database (HMDB) | |
| URL | http://www.hmdb.ca/metabolites/HMDB0015048 | |
| Description | The Human Metabolome Database (HMDB) is a freely available electronic database containing detailed information about small molecule metabolites found in the human body. | |
| Explanation | HMDB is offered to the public as a freely available resource. Use and re-distribution of the data, in whole or in part, for commercial purposes requires explicit permission of the authors and explicit acknowledgment of the source material (HMDB) and the original publication (see the HMDB citing page). We ask that users who download significant portions of the database cite the HMDB paper in any resulting publications. | |
Solubility |
>67.9 [ug/mL] (The mean of the results at pH 7.4), 2.94e-03 g/L | |
| Record name | SID49648522 | |
| Source | Burnham Center for Chemical Genomics | |
| URL | https://pubchem.ncbi.nlm.nih.gov/bioassay/1996#section=Data-Table | |
| Description | Aqueous solubility in buffer at pH 7.4 | |
| Record name | Repaglinide | |
| Source | Human Metabolome Database (HMDB) | |
| URL | http://www.hmdb.ca/metabolites/HMDB0015048 | |
| Description | The Human Metabolome Database (HMDB) is a freely available electronic database containing detailed information about small molecule metabolites found in the human body. | |
| Explanation | HMDB is offered to the public as a freely available resource. Use and re-distribution of the data, in whole or in part, for commercial purposes requires explicit permission of the authors and explicit acknowledgment of the source material (HMDB) and the original publication (see the HMDB citing page). We ask that users who download significant portions of the database cite the HMDB paper in any resulting publications. | |
Mechanism of Action |
Repaglinide activity is dependent on the presence functioning β cells and glucose. In contrast to sulfonylurea insulin secretatogogues, repaglinide has no effect on insulin release in the absence of glucose. Rather, it potentiates the effect of extracellular glucose on ATP-sensitive potassium channel and has little effect on insulin levels between meals and overnight. As such, repaglinide is more effective at reducing postprandial blood glucose levels than fasting blood glucose levels and requires a longer duration of therapy (approximately one month) before decreases in fasting blood glucose are observed. The insulinotropic effects of repaglinide are highest at intermediate glucose levels (3 to 10 mmol/L) and it does not increase insulin release already stimulated by high glucose concentrations (greater than 15 mmol/L). Repaglinide appears to be selective for pancreatic β cells and does not appear to affect skeletal or cardiac muscle or thyroid tissue. | |
| Record name | Repaglinide | |
| Source | DrugBank | |
| URL | https://www.drugbank.ca/drugs/DB00912 | |
| Description | The DrugBank database is a unique bioinformatics and cheminformatics resource that combines detailed drug (i.e. chemical, pharmacological and pharmaceutical) data with comprehensive drug target (i.e. sequence, structure, and pathway) information. | |
| Explanation | Creative Common's Attribution-NonCommercial 4.0 International License (http://creativecommons.org/licenses/by-nc/4.0/legalcode) | |
CAS No. |
135062-02-1 | |
| Record name | Repaglinide | |
| Source | CAS Common Chemistry | |
| URL | https://commonchemistry.cas.org/detail?cas_rn=135062-02-1 | |
| Description | CAS Common Chemistry is an open community resource for accessing chemical information. Nearly 500,000 chemical substances from CAS REGISTRY cover areas of community interest, including common and frequently regulated chemicals, and those relevant to high school and undergraduate chemistry classes. This chemical information, curated by our expert scientists, is provided in alignment with our mission as a division of the American Chemical Society. | |
| Explanation | The data from CAS Common Chemistry is provided under a CC-BY-NC 4.0 license, unless otherwise stated. | |
| Record name | Repaglinide [USAN:USP:INN:BAN] | |
| Source | ChemIDplus | |
| URL | https://pubchem.ncbi.nlm.nih.gov/substance/?source=chemidplus&sourceid=0135062021 | |
| Description | ChemIDplus is a free, web search system that provides access to the structure and nomenclature authority files used for the identification of chemical substances cited in National Library of Medicine (NLM) databases, including the TOXNET system. | |
| Record name | Repaglinide | |
| Source | DrugBank | |
| URL | https://www.drugbank.ca/drugs/DB00912 | |
| Description | The DrugBank database is a unique bioinformatics and cheminformatics resource that combines detailed drug (i.e. chemical, pharmacological and pharmaceutical) data with comprehensive drug target (i.e. sequence, structure, and pathway) information. | |
| Explanation | Creative Common's Attribution-NonCommercial 4.0 International License (http://creativecommons.org/licenses/by-nc/4.0/legalcode) | |
| Record name | Repaglinide | |
| Source | DTP/NCI | |
| URL | https://dtp.cancer.gov/dtpstandard/servlet/dwindex?searchtype=NSC&outputformat=html&searchlist=759893 | |
| Description | The NCI Development Therapeutics Program (DTP) provides services and resources to the academic and private-sector research communities worldwide to facilitate the discovery and development of new cancer therapeutic agents. | |
| Explanation | Unless otherwise indicated, all text within NCI products is free of copyright and may be reused without our permission. Credit the National Cancer Institute as the source. | |
| Record name | Repaglinide | |
| Source | EPA DSSTox | |
| URL | https://comptox.epa.gov/dashboard/DTXSID3023552 | |
| Description | DSSTox provides a high quality public chemistry resource for supporting improved predictive toxicology. | |
| Record name | (+)-2-Ethoxy-alpha-(((S)-alpha-isobutyl-o-piperidinobenzyl)carbamoyl)-p-toluic acid | |
| Source | European Chemicals Agency (ECHA) | |
| URL | https://echa.europa.eu/information-on-chemicals | |
| Description | The European Chemicals Agency (ECHA) is an agency of the European Union which is the driving force among regulatory authorities in implementing the EU's groundbreaking chemicals legislation for the benefit of human health and the environment as well as for innovation and competitiveness. | |
| Explanation | Use of the information, documents and data from the ECHA website is subject to the terms and conditions of this Legal Notice, and subject to other binding limitations provided for under applicable law, the information, documents and data made available on the ECHA website may be reproduced, distributed and/or used, totally or in part, for non-commercial purposes provided that ECHA is acknowledged as the source: "Source: European Chemicals Agency, http://echa.europa.eu/". Such acknowledgement must be included in each copy of the material. ECHA permits and encourages organisations and individuals to create links to the ECHA website under the following cumulative conditions: Links can only be made to webpages that provide a link to the Legal Notice page. | |
| Record name | REPAGLINIDE | |
| Source | FDA Global Substance Registration System (GSRS) | |
| URL | https://gsrs.ncats.nih.gov/ginas/app/beta/substances/668Z8C33LU | |
| Description | The FDA Global Substance Registration System (GSRS) enables the efficient and accurate exchange of information on what substances are in regulated products. Instead of relying on names, which vary across regulatory domains, countries, and regions, the GSRS knowledge base makes it possible for substances to be defined by standardized, scientific descriptions. | |
| Explanation | Unless otherwise noted, the contents of the FDA website (www.fda.gov), both text and graphics, are not copyrighted. They are in the public domain and may be republished, reprinted and otherwise used freely by anyone without the need to obtain permission from FDA. Credit to the U.S. Food and Drug Administration as the source is appreciated but not required. | |
| Record name | Repaglinide | |
| Source | Human Metabolome Database (HMDB) | |
| URL | http://www.hmdb.ca/metabolites/HMDB0015048 | |
| Description | The Human Metabolome Database (HMDB) is a freely available electronic database containing detailed information about small molecule metabolites found in the human body. | |
| Explanation | HMDB is offered to the public as a freely available resource. Use and re-distribution of the data, in whole or in part, for commercial purposes requires explicit permission of the authors and explicit acknowledgment of the source material (HMDB) and the original publication (see the HMDB citing page). We ask that users who download significant portions of the database cite the HMDB paper in any resulting publications. | |
Melting Point |
130-131 °C, 130 - 131 °C | |
| Record name | Repaglinide | |
| Source | DrugBank | |
| URL | https://www.drugbank.ca/drugs/DB00912 | |
| Description | The DrugBank database is a unique bioinformatics and cheminformatics resource that combines detailed drug (i.e. chemical, pharmacological and pharmaceutical) data with comprehensive drug target (i.e. sequence, structure, and pathway) information. | |
| Explanation | Creative Common's Attribution-NonCommercial 4.0 International License (http://creativecommons.org/licenses/by-nc/4.0/legalcode) | |
| Record name | Repaglinide | |
| Source | Human Metabolome Database (HMDB) | |
| URL | http://www.hmdb.ca/metabolites/HMDB0015048 | |
| Description | The Human Metabolome Database (HMDB) is a freely available electronic database containing detailed information about small molecule metabolites found in the human body. | |
| Explanation | HMDB is offered to the public as a freely available resource. Use and re-distribution of the data, in whole or in part, for commercial purposes requires explicit permission of the authors and explicit acknowledgment of the source material (HMDB) and the original publication (see the HMDB citing page). We ask that users who download significant portions of the database cite the HMDB paper in any resulting publications. | |
Synthesis routes and methods
Procedure details






Retrosynthesis Analysis
AI-Powered Synthesis Planning: Our tool employs the Template_relevance Pistachio, Template_relevance Bkms_metabolic, Template_relevance Pistachio_ringbreaker, Template_relevance Reaxys, Template_relevance Reaxys_biocatalysis model, leveraging a vast database of chemical reactions to predict feasible synthetic routes.
One-Step Synthesis Focus: Specifically designed for one-step synthesis, it provides concise and direct routes for your target compounds, streamlining the synthesis process.
Accurate Predictions: Utilizing the extensive PISTACHIO, BKMS_METABOLIC, PISTACHIO_RINGBREAKER, REAXYS, REAXYS_BIOCATALYSIS database, our tool offers high-accuracy predictions, reflecting the latest in chemical research and data.
Strategy Settings
| Precursor scoring | Relevance Heuristic |
|---|---|
| Min. plausibility | 0.01 |
| Model | Template_relevance |
| Template Set | Pistachio/Bkms_metabolic/Pistachio_ringbreaker/Reaxys/Reaxys_biocatalysis |
| Top-N result to add to graph | 6 |
Feasible Synthetic Routes
体外研究产品的免责声明和信息
请注意,BenchChem 上展示的所有文章和产品信息仅供信息参考。 BenchChem 上可购买的产品专为体外研究设计,这些研究在生物体外进行。体外研究,源自拉丁语 "in glass",涉及在受控实验室环境中使用细胞或组织进行的实验。重要的是要注意,这些产品没有被归类为药物或药品,他们没有得到 FDA 的批准,用于预防、治疗或治愈任何医疗状况、疾病或疾病。我们必须强调,将这些产品以任何形式引入人类或动物的身体都是法律严格禁止的。遵守这些指南对确保研究和实验的法律和道德标准的符合性至关重要。